Explaining protein–protein interactions with knowledge graph-based semantic similarity
- Conference/Journal: Computers in Biology and Medicine
- Year: 2024
- Full paper URL: kgsim2vec
Abstract
The application of artificial intelligence and machine learning methods for several biomedical applications, such as protein–protein interaction prediction, has gained significant traction in recent decades. However, explainability is a key aspect of using machine learning as a tool for scientific discovery. Explainable artificial intelligence approaches help clarify algorithmic mechanisms and identify potential bias in the data.
Given the complexity of the biomedical domain, explanations should be grounded in domain knowledge which can be achieved by using ontologies and knowledge graphs. These knowledge graphs express knowledge about a domain by capturing different perspectives of the representation of real-world entities. However, the most popular way to explore knowledge graphs with machine learning is through using embeddings, which are not explainable. As an alternative, knowledge graph-based semantic similarity offers the advantage of being explainable. Additionally, similarity can be computed to capture different semantic aspects within the knowledge graph and increasing the explainability of predictive approaches.
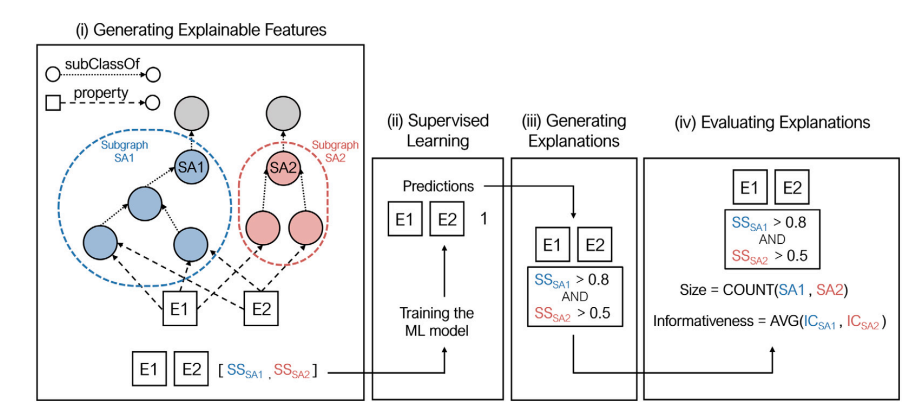
We propose a novel method to generate explainable vector representations, KGsim2vec, that uses aspectoriented semantic similarity features to represent pairs of entities in a knowledge graph. Our approach employs a set of machine learning models, including decision trees, genetic programming, random forest and eXtreme gradient boosting, to predict relations between entities.
The experiments reveal that considering multiple semantic aspects when representing the similarity between two entities improves explainability and predictive performance. KGsim2vec performs better than black-box methods based on knowledge graph embeddings or graph neural networks. Moreover, KGsim2vec produces global models that can capture biological phenomena and elucidate data biases.